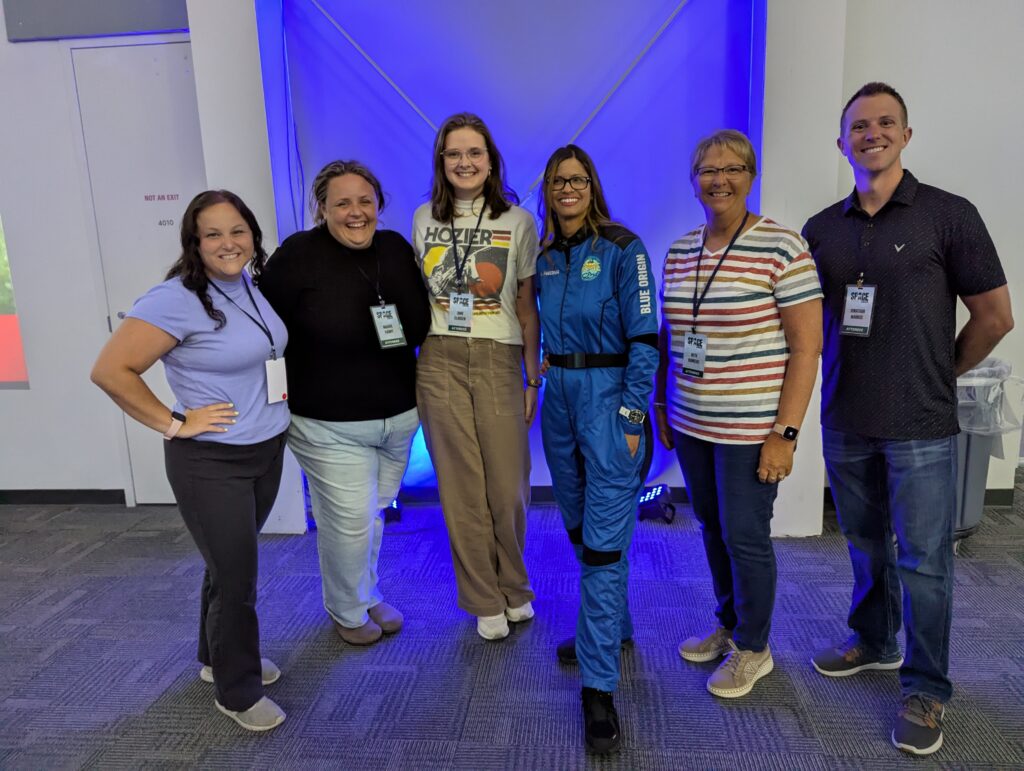
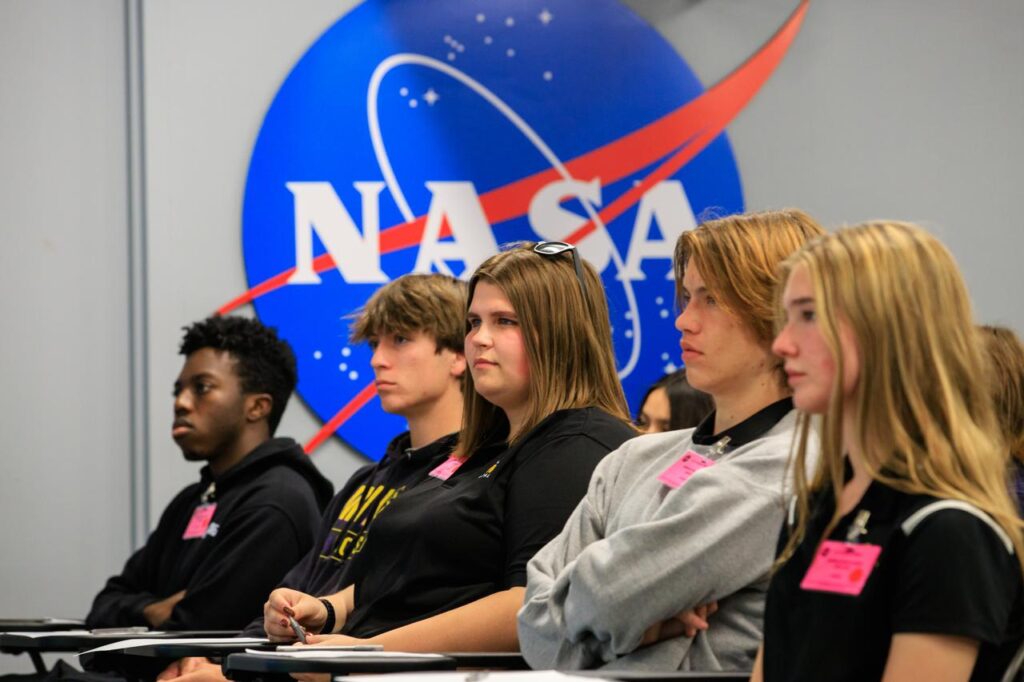
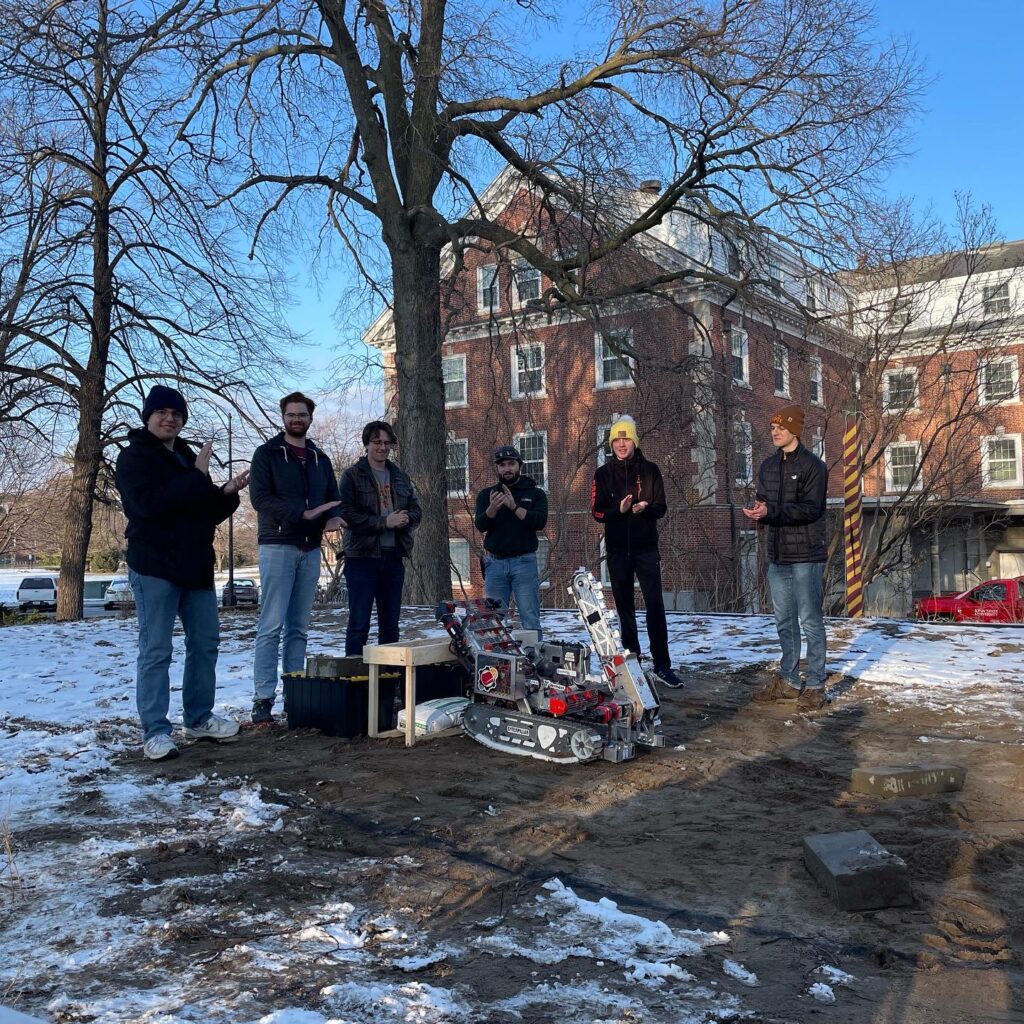
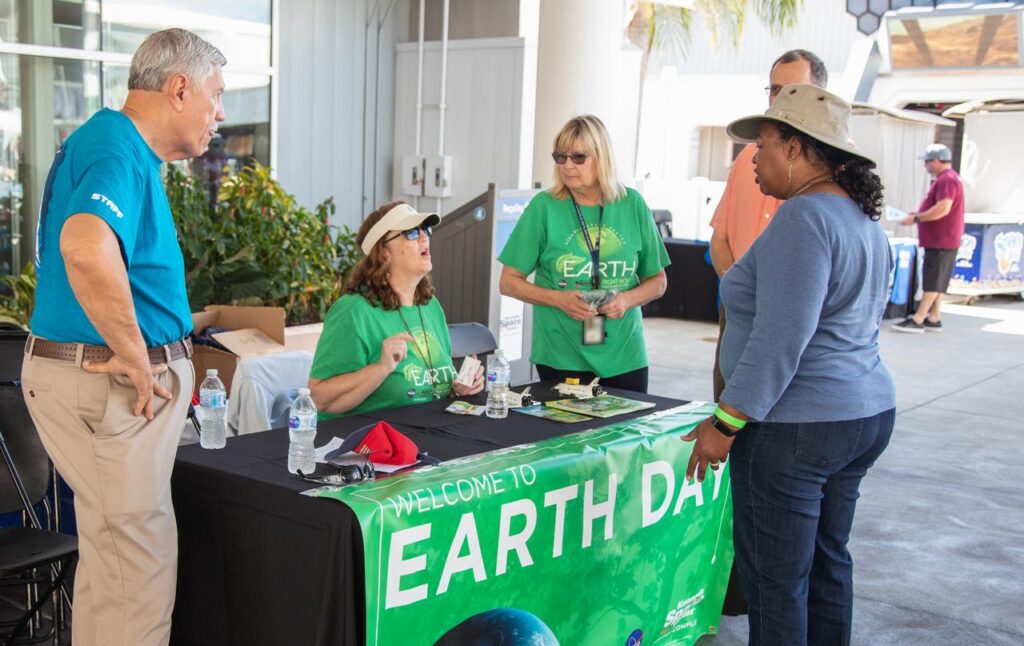
🚀 From Classroom to Cosmos: Iowa Educators Soar at SPACE 2025
A team of Iowa educators and STEM leaders recently experienced a once-in-a-lifetime opportunity at the SPACE 2025 Port Area Conference for Educators, hosted at the Kennedy Space Center Visitor Complex